Advertisements
Advertisements
प्रश्न
Which of the following enzymes catalyse the removal of nucleotides from the ends of DNA?
पर्याय
Endonuclease
Exonuclease
DNA ligase
Hind - II
Advertisements
उत्तर
Exonuclease
Explanation:
The enzyme that catalyzes the removal of nucleotides from the ends of DNA is exonuclease. Exonuclease specifically removes nucleotides one by one from the ends of a DNA strand, whereas endonucleases cut DNA at internal sites. DNA ligase joins DNA fragments but does not remove nucleotides. Hind II is a type of restriction endonuclease that cuts DNA at specific internal sequences, not at the ends.
संबंधित प्रश्न
Why is the enzyme cellulase needed for isolating genetic material from plant cells and not form the animal cells?
How does a restriction nuclease function? Explain
Do eukaryotic cells have restriction endonucleases? Justify your answer.
Collect 5 examples of palindromic DNA sequences. Better try to create a palindromic sequence by following base-pair rules.
Distinguish between exonuclease and endonuclease.
How does restriction endonuclease function?
Answer the following question.
Explain the significance of palindromic nucleotide sequence in the formation of recombinant DNA.
Answer the following question.
Write the use of restriction endonuclease in the formation of recombinant DNA.
Give a reason why :
Single cloning site is preferred in a vector.
A mixture containing DNA fragments a, b, c and d, with molecular weights of a + b = c, a > b and d > c was subject to agarose get electrophoresis. This position of these fragments from cathode to anode to anode sides of the gel would be ______.
Which of the following radioisotope is not suitable for DNA labeling based studies?
Restriction enzymes ______.
Molecular scissors, which cut DNA at specific site is ______.
Which of the given statements is correct in the context of visualizing DNA molecules separated by agarose gel electrophoresis?
While isolating DNA from bacteria, which of the following enzymes is not required?
Would you choose an exonuclease while producing a recombinant DNA molecule?
Restriction enzymes should not have more than one site of action in the cloning site of a vector. Comment.
How does one visualise DNA on an agarose gel?
A mixture of fragmented DNA was electrophoresed in an agarose gel. After staining the gel with ethidium bromide, no DNA bands were observed. What could be the reason?
CTTAAG
GAATTC
- What are such sequences called? Name the enzyme used that recognizes such nucleotide sequences.
- What is their significance in biotechnology?
Given below is the stepwise schematic representation of the process of electrophoresis. Identify the 'alphabets' representing
- Anode end
- smallest/lightest DNA strand in the matrix
- Agarose gel
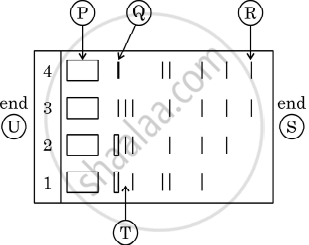
What is elution?
Given below is the restriction site of a restriction endonuclease Pst-I and the cleavage sites on a DNA molecule.
\[\ce{5' C - T - G - C - A \overset{\downarrow}{-}{G 3'}}\]
\[\ce{3' G\underset{\uparrow}{-} A - C - G - T - C 5'}\]
Choose the option that gives the correct resultant fragments by the action of the enzyme Pst-I.
What are the protruding and hanging stretches of DNA produced by these restriction enzymes called? Describe their role in the formation of rDNA.
