Advertisements
Advertisements
प्रश्न
Which of the following bacteria is not a source of restriction endonuclease?
विकल्प
Haemophilus influenzae
Escherichia coli
Entamoeba coli
Bacillus amyloliquefaciens
Advertisements
उत्तर
Entamoeba coli
APPEARS IN
संबंधित प्रश्न
Explain with the help of a suitable example the naming of a restriction endonuclease.
How are 'sticky ends' formed on a DNA strand? Why are they so called?
Mention the difference in the mode of action of exonuclease and endonuclease.
Name the enzymes that are used for the isolation of DNA from bacterial and fungal cells for recombinant DNA technology.
Collect 5 examples of palindromic DNA sequences. Better try to create a palindromic sequence by following base-pair rules.
Distinguish between exonuclease and endonuclease.
Answer the following question.
Write the use of restriction endonuclease in the formation of recombinant DNA.
Molecular scissors, which cut DNA at specific site is ______.
DNA fragments separate according to size through?
A specific recognition sequence identified by endonucleases to make cuts at specific positions within the DNA is ______
Which of the following enzymes catalyse the removal of nucleotides from the ends of DNA?
Which of the given statements is correct in the context of visualizing DNA molecules separated by agarose gel electrophoresis?
In agarose gel electrophoresis, DNA molecules are separated on the basis of their ______.
Restriction enzymes that are used in the construction of recombinant DNA are endonucleases which cut the DNA at ‘specific-recognition sequence’. What would be the disadvantage if they do not cut the DNA at specific-recognition sequence?
How does one visualise DNA on an agarose gel?
CTTAAG
GAATTC
- What are such sequences called? Name the enzyme used that recognizes such nucleotide sequences.
- What is their significance in biotechnology?
Given below is the stepwise schematic representation of the process of electrophoresis. Identify the 'alphabets' representing
- Anode end
- smallest/lightest DNA strand in the matrix
- Agarose gel
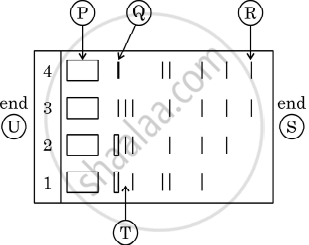
State the principle involved in separation of DNA fragments using gel electrophoresis.
How are DNA fragments visualised once they are separated by gel electrophoresis?
