Advertisements
Advertisements
प्रश्न
Given below is the restriction site of a restriction endonuclease Pst-I and the cleavage sites on a DNA molecule.
\[\ce{5' C - T - G - C - A \overset{\downarrow}{-}{G 3'}}\]
\[\ce{3' G\underset{\uparrow}{-} A - C - G - T - C 5'}\]
Choose the option that gives the correct resultant fragments by the action of the enzyme Pst-I.
विकल्प
5' C - T - G C - A - G 3' 3' G - A - C - G - T C 5' 5' C - T G - C - A - G 3' 3' G - A - G - C T - C 5' 5' C - T - G - C A - G 3' 3' G - A - C - G T - C 5' 5' C - T - G - C - A G 3' 3' G A - C - G - T - C 5'
Advertisements
उत्तर
| 5' C - T - G - C - A | G 3' |
| 3' G | A - C - G - T - C 5' |
Explanation:
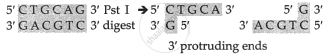
Enzyme Pst-I makes a staggered cut of the DNA at its recognition sequence.
संबंधित प्रश्न
Explain briefly:
Restriction enzymes and DNA
Distinguish between exonuclease and endonuclease.
Give a reason why :
Single cloning site is preferred in a vector.
A mixture containing DNA fragments a, b, c and d, with molecular weights of a + b = c, a > b and d > c was subject to agarose get electrophoresis. This position of these fragments from cathode to anode to anode sides of the gel would be ______.
What does H in’ ‘d’ and ‘III’ refer to in the enzyme Hind III?
Restriction enzymes should not have more than one site of action in the cloning site of a vector. Comment.
How does one visualise DNA on an agarose gel?
CTTAAG
GAATTC
- What are such sequences called? Name the enzyme used that recognizes such nucleotide sequences.
- What is their significance in biotechnology?
What is elution?
State the principle involved in separation of DNA fragments using gel electrophoresis.
