Advertisements
Advertisements
प्रश्न
Which of the following bacteria is not a source of restriction endonuclease?
पर्याय
Haemophilus influenzae
Escherichia coli
Entamoeba coli
Bacillus amyloliquefaciens
Advertisements
उत्तर
Entamoeba coli
APPEARS IN
संबंधित प्रश्न
Explain with the help of a suitable example the naming of a restriction endonuclease.
Mention the difference in the mode of action of exonuclease and endonuclease.
Name and describe the technique that helps in separating the DNA fragments formed by the use of restriction endonuclease
Distinguish between exonuclease and endonuclease.
Give a reason why :
Single cloning site is preferred in a vector.
The total number of nucleotide sequences of DNA that code for a hormone is 1530. The proportion of different bases in the sequence is found to be Adenine = 34%, Guanine = 19%, Cytosine = 23%, Thymine = 19%.
Applying Chargaff’s rule, what conclusion can be drawn?
There is a restriction endonudease called as EcoRI. What does co part in it stands for?
Which of the following enzymes catalyse the removal of nucleotides from the ends of DNA?
In agarose gel electrophoresis, DNA molecules are separated on the basis of their ______.
The role of DNA ligase in the construction of a recombinant DNA molecule is ______.
Which of the following statements does not hold true for restriction enzyme?
What does H in’ ‘d’ and ‘III’ refer to in the enzyme Hind III?
How does one visualise DNA on an agarose gel?
Carefully observe the given picture. A mixture of DNA with fragments ranging from 200 base pairs to 2500 base pairs was electrophoresed on agarose gel with the following arrangement.
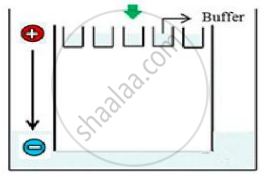
(a) What result will be obtained on staining with ethidium bromide? Explain with reason.
(b) The above setup was modified and a band with 250 base pairs was obtained at X.
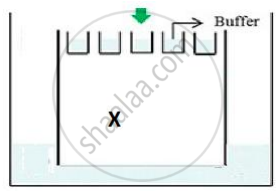
What change(s) were made to the previous design to obtain a band at X? Why did the band appear at position X?
What is elution?
State the principle involved in separation of DNA fragments using gel electrophoresis.
