Advertisements
Advertisements
प्रश्न
Carefully observe the given picture. A mixture of DNA with fragments ranging from 200 base pairs to 2500 base pairs was electrophoresed on agarose gel with the following arrangement.
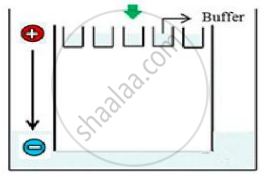
(a) What result will be obtained on staining with ethidium bromide? Explain with reason.
(b) The above setup was modified and a band with 250 base pairs was obtained at X.
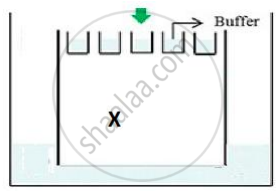
What change(s) were made to the previous design to obtain a band at X? Why did the band appear at position X?
Advertisements
उत्तर
(a) No bands will be obtained as all DNA will be seen in the well only; DNA fragments being negatively charged will not move towards the negative end/cathode. DNA being negatively charged will remain stationed at the positive end/anode end of the agar block.
(b)
- The position of the positive terminal/end/anode and the negative terminal/end/cathode was inter-changed.
- The fragment with the least base pairs will get separated faster and move faster to the anode end.
APPEARS IN
संबंधित प्रश्न
How are 'sticky ends' formed on a DNA strand? Why are they so called?
Suggest a technique to a researcher who needs to separate fragments of DNA.
Name the enzymes that are used for the isolation of DNA from bacterial and fungal cells for recombinant DNA technology.
How does a restriction nuclease function? Explain
Make a chart (with diagrammatic representation) showing a restriction enzyme, the substrate DNA on which it acts, the site at which it cuts DNA and the product it produces.
Give a reason why :
Single cloning site is preferred in a vector.
Which of the following statements does not hold true for restriction enzyme?
State the principle involved in separation of DNA fragments using gel electrophoresis.
How are DNA fragments visualised once they are separated by gel electrophoresis?
Hind II always cuts DNA molecules at a particular point called recognition sequence and it consists of ______.
