Advertisements
Advertisements
Question
Explain briefly:
Restriction enzymes and DNA
Advertisements
Solution
Restriction enzymes belong to a larger class of enzymes called nucleases. They are mainly classified into two types:
- Exonucleases: They remove nucleotides from the terminal ends of the DNA molecule.
- Endonucleases: They make cuts at specific positions anywhere within the DNA molecule.
Restriction endonucleases are a specific class of endonucleases that recognise palindromic nucleotide sequences and cleave DNA at specific sites. They cut both strands of the DNA double helix at specific points to generate sticky or blunt ends, which are essential for creating recombinant DNA.
APPEARS IN
RELATED QUESTIONS
How are 'sticky ends' formed on a DNA strand? Why are they so called?
Why is the enzyme cellulase needed for isolating genetic material from plant cells and not form the animal cells?
Mention the difference in the mode of action of exonuclease and endonuclease.
How does a restriction nuclease function? Explain
Make a chart (with diagrammatic representation) showing a restriction enzyme, the substrate DNA on which it acts, the site at which it cuts DNA and the product it produces.
Answer the following question.
Write the use of restriction endonuclease in the formation of recombinant DNA.
Give a reason why :
Single cloning site is preferred in a vector.
The total number of nucleotide sequences of DNA that code for a hormone is 1530. The proportion of different bases in the sequence is found to be Adenine = 34%, Guanine = 19%, Cytosine = 23%, Thymine = 19%.
Applying Chargaff’s rule, what conclusion can be drawn?
The DNA fragment separated on an agarose gel can be visualized by staining with ______.
DNA fragments separate according to size through?
DNA strands on a gel stained with ethidium bromide when viewed under UV radiation, appear as ______
'Restriction' in Restriction enzyme refers to ______.
In agarose gel electrophoresis, DNA molecules are separated on the basis of their ______.
While isolating DNA from bacteria, which of the following enzymes is not required?
Which of the following bacteria is not a source of restriction endonuclease?
Which of the following statements does not hold true for restriction enzyme?
What does H in’ ‘d’ and ‘III’ refer to in the enzyme Hind III?
CTTAAG
GAATTC
- What are such sequences called? Name the enzyme used that recognizes such nucleotide sequences.
- What is their significance in biotechnology?
Given below is the stepwise schematic representation of the process of electrophoresis. Identify the 'alphabets' representing
- Anode end
- smallest/lightest DNA strand in the matrix
- Agarose gel
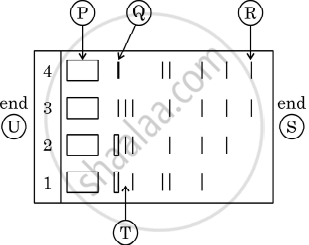
What is elution?
'EcoRI' has played a very significant role in rDNA technology.
- Explain the convention for naming EcoRI.
- Write the recognition site and the cleavage sites of this restriction endonuclease.
State the principle involved in separation of DNA fragments using gel electrophoresis.
Identify the activity of endonuclease and exonuclease in the given image.
