Advertisements
Advertisements
Question
'EcoRI' has played a very significant role in rDNA technology.
- Explain the convention for naming EcoRI.
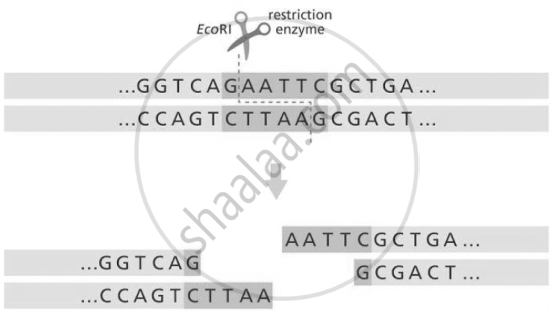
- Write the recognition site and the cleavage sites of this restriction endonuclease.
Advertisements
Solution
(I) Restriction endonucleases are named as follows:
1st alphabet represents the genus of the organism from which the enzyme is isolated.
2nd and 3rd alphabet represent the species of the organism.
4th alphabet represents the strain.
The Roman number represents the order of isolation or discovery of the enzyme.
EcoRI comes from the Escherichia coli RYB strain.
In EcoRI, 'E' comes from the genus 'Escherichia' and 'co' comes from the species name 'Coli'.
The letter 'R' is derived from the name of the strain RYB.
Roman numbers following the names indicate the order in which the enzymes were isolated from that strain of bacteria.
(II) The recognition sequence where EcoRI cleaves the DNA molecule is G/AATTC. Such sequences have a complementary sequence, CTTAA/G, which is known as a palindromic sequence.
APPEARS IN
RELATED QUESTIONS
How are 'sticky ends' formed on a DNA strand? Why are they so called?
How does a restriction nuclease function? Explain
Name and describe the technique that helps in separating the DNA fragments formed by the use of restriction endonuclease
Make a chart (with diagrammatic representation) showing a restriction enzyme, the substrate DNA on which it acts, the site at which it cuts DNA and the product it produces.
The total number of nucleotide sequences of DNA that code for a hormone is 1530. The proportion of different bases in the sequence is found to be Adenine = 34%, Guanine = 19%, Cytosine = 23%, Thymine = 19%.
Applying Chargaff’s rule, what conclusion can be drawn?
Which of the given statements is correct in the context of visualizing DNA molecules separated by agarose gel electrophoresis?
Which of the following bacteria is not a source of restriction endonuclease?
Would you choose an exonuclease while producing a recombinant DNA molecule?
How does one visualise DNA on an agarose gel?
What is elution?
