Advertisements
Advertisements
Questions
Distinguish between exonuclease and endonuclease.
Differentiate between exonuclease and endonuclease.
Discuss with your teacher and find out how to distinguish between exonuclease and endonuclease.
Advertisements
Solution 1
| Sr. No. | Exonuclease | Endonuclease |
| 1. | It is a type of restriction enzyme that removes the nucleotide from the 5' or 3' ends of the DNA molecule. | It is a type of restriction enzyme that makes a cut within the DNA at a specific site to generate sticky ends. |
| 2. | They separated DNA base pairs at their terminal ends.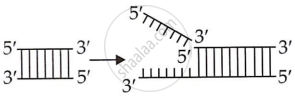 |
They separate DNA at any location other than the terminal ends.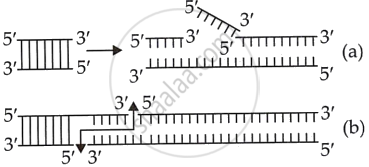 |
| 3. | They work on single strands of DNA or gaps in double-stranded DNA. | They cut one or both strands of double-stranded DNA. |
Solution 2
| Feature | Exonuclease | Endonuclease |
| Clevage Site: | Eliminates nucleotides from DNA/RNA ends. | Cuts DNA within the strand at specific sites. |
| Requirement of Free Ends: | Needs free 3' or 5' ends (in one duplex strand). | Requires just one or both strands of the DNA duplex and doesn’t require free ends. |
| Mode of Action: | Progressively removes nucleotides one by one. | Cuts at specific recognition sequences. |
| Example: | DNA polymerase I (has exonuclease activity). | EcoRI, HindIII, BamHI (restriction enzymes). |
RELATED QUESTIONS
Why is the enzyme cellulase needed for isolating genetic material from plant cells and not form the animal cells?
Suggest a technique to a researcher who needs to separate fragments of DNA.
Name the enzymes that are used for the isolation of DNA from bacterial and fungal cells for recombinant DNA technology.
Make a chart (with diagrammatic representation) showing a restriction enzyme, the substrate DNA on which it acts, the site at which it cuts DNA and the product it produces.
Collect 5 examples of palindromic DNA sequences. Better try to create a palindromic sequence by following base-pair rules.
Explain briefly:
Restriction enzymes and DNA
Explain the roles of the following with the help of an example each in recombinant DNA technology :
Restriction Enzymes
Answer the following question.
Explain the significance of palindromic nucleotide sequence in the formation of recombinant DNA.
The total number of nucleotide sequences of DNA that code for a hormone is 1530. The proportion of different bases in the sequence is found to be Adenine = 34%, Guanine = 19%, Cytosine = 23%, Thymine = 19%.
Applying Chargaff’s rule, what conclusion can be drawn?
Which of the following radioisotope is not suitable for DNA labeling based studies?
There is a restriction endonudease called as EcoRI. What does co part in it stands for?
'Restriction' in restriction enzyme refers to
A specific recognition sequence identified by endonucleases to make cuts at specific positions within the DNA is ______
Which of the following enzymes catalyse the removal of nucleotides from the ends of DNA?
'Restriction' in Restriction enzyme refers to ______.
Which of the following bacteria is not a source of restriction endonuclease?
Which of the following statements does not hold true for restriction enzyme?
Restriction enzymes should not have more than one site of action in the cloning site of a vector. Comment.
A plasmid DNA and a linear DNA (both are of the same size) have one site for a restriction endonuclease. When cut and separated on agarose gel electrophoresis, plasmid shows one DNA band while linear DNA shows two fragments. Explain.
How does one visualise DNA on an agarose gel?
Carefully observe the given picture. A mixture of DNA with fragments ranging from 200 base pairs to 2500 base pairs was electrophoresed on agarose gel with the following arrangement.
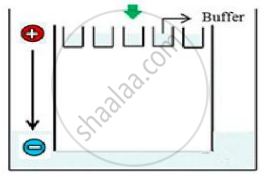
(a) What result will be obtained on staining with ethidium bromide? Explain with reason.
(b) The above setup was modified and a band with 250 base pairs was obtained at X.
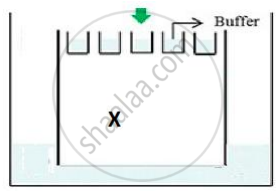
What change(s) were made to the previous design to obtain a band at X? Why did the band appear at position X?
Given below is the stepwise schematic representation of the process of electrophoresis. Identify the 'alphabets' representing
- Anode end
- smallest/lightest DNA strand in the matrix
- Agarose gel
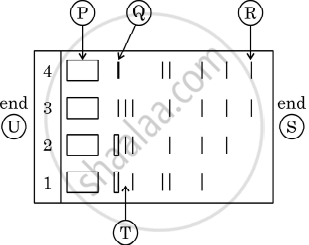
State the importance of elution in this process.
What are the protruding and hanging stretches of DNA produced by these restriction enzymes called? Describe their role in the formation of rDNA.
Hind II always cuts DNA molecules at a particular point called recognition sequence and it consists of ______.
