Advertisements
Advertisements
Question
A plasmid DNA and a linear DNA (both are of the same size) have one site for a restriction endonuclease. When cut and separated on agarose gel electrophoresis, plasmid shows one DNA band while linear DNA shows two fragments. Explain.
Advertisements
Solution
It is because plasmid is a circular DNA molecule. When cut with enzyme, it becomes linear but does not get fragmented. Whereas, a linear DNA molecule gets cut into two fragments. Hence, a single DNA band is observed for plasmid while two DNA bands are observed for linear DNA in agarose gel.

APPEARS IN
RELATED QUESTIONS
How are 'sticky ends' formed on a DNA strand? Why are they so called?
Mention the difference in the mode of action of exonuclease and endonuclease.
Do eukaryotic cells have restriction endonucleases? Justify your answer.
Distinguish between exonuclease and endonuclease.
How does restriction endonuclease function?
Explain the roles of the following with the help of an example each in recombinant DNA technology :
Restriction Enzymes
A mixture containing DNA fragments a, b, c and d, with molecular weights of a + b = c, a > b and d > c was subject to agarose get electrophoresis. This position of these fragments from cathode to anode to anode sides of the gel would be ______.
Which of the following radioisotope is not suitable for DNA labeling based studies?
Molecular scissors, which cut DNA at specific site is ______.
Which of the given statements is correct in the context of visualizing DNA molecules separated by agarose gel electrophoresis?
While isolating DNA from bacteria, which of the following enzymes is not required?
Which of the following bacteria is not a source of restriction endonuclease?
Restriction enzymes should not have more than one site of action in the cloning site of a vector. Comment.
Restriction enzymes that are used in the construction of recombinant DNA are endonucleases which cut the DNA at ‘specific-recognition sequence’. What would be the disadvantage if they do not cut the DNA at specific-recognition sequence?
How does one visualise DNA on an agarose gel?
Carefully observe the given picture. A mixture of DNA with fragments ranging from 200 base pairs to 2500 base pairs was electrophoresed on agarose gel with the following arrangement.
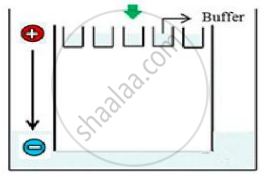
(a) What result will be obtained on staining with ethidium bromide? Explain with reason.
(b) The above setup was modified and a band with 250 base pairs was obtained at X.
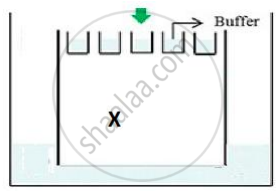
What change(s) were made to the previous design to obtain a band at X? Why did the band appear at position X?
Given below is the restriction site of a restriction endonuclease Pst-I and the cleavage sites on a DNA molecule.
\[\ce{5' C - T - G - C - A \overset{\downarrow}{-}{G 3'}}\]
\[\ce{3' G\underset{\uparrow}{-} A - C - G - T - C 5'}\]
Choose the option that gives the correct resultant fragments by the action of the enzyme Pst-I.
State the principle involved in separation of DNA fragments using gel electrophoresis.
