Advertisements
Advertisements
प्रश्न
Restriction enzymes that are used in the construction of recombinant DNA are endonucleases which cut the DNA at ‘specific-recognition sequence’. What would be the disadvantage if they do not cut the DNA at specific-recognition sequence?
Advertisements
उत्तर
If the restriction enzymes would cut DNA at random sites instead of at specific sites, then the DNA fragments obtained will not have ‘sticky ends’. In the absence of sticky ends, construction of recombinant DNA molecules would not be possible.
APPEARS IN
संबंधित प्रश्न
Explain with the help of a suitable example the naming of a restriction endonuclease.
How are 'sticky ends' formed on a DNA strand? Why are they so called?
Name the enzymes that are used for the isolation of DNA from bacterial and fungal cells for recombinant DNA technology.
How does a restriction nuclease function? Explain
Name and describe the technique that helps in separating the DNA fragments formed by the use of restriction endonuclease
Make a chart (with diagrammatic representation) showing a restriction enzyme, the substrate DNA on which it acts, the site at which it cuts DNA and the product it produces.
Answer the following question.
Explain the significance of palindromic nucleotide sequence in the formation of recombinant DNA.
Give a reason why :
Single cloning site is preferred in a vector.
Restriction enzymes ______.
Which of the following enzymes catalyse the removal of nucleotides from the ends of DNA?
'Restriction' in Restriction enzyme refers to ______.
While isolating DNA from bacteria, which of the following enzymes is not required?
Would you choose an exonuclease while producing a recombinant DNA molecule?
Restriction enzymes should not have more than one site of action in the cloning site of a vector. Comment.
A plasmid DNA and a linear DNA (both are of the same size) have one site for a restriction endonuclease. When cut and separated on agarose gel electrophoresis, plasmid shows one DNA band while linear DNA shows two fragments. Explain.
CTTAAG
GAATTC
- What are such sequences called? Name the enzyme used that recognizes such nucleotide sequences.
- What is their significance in biotechnology?
Given below is the stepwise schematic representation of the process of electrophoresis. Identify the 'alphabets' representing
- Anode end
- smallest/lightest DNA strand in the matrix
- Agarose gel
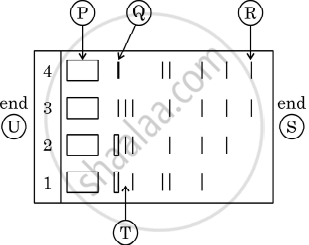
State the importance of elution in this process.
What are the protruding and hanging stretches of DNA produced by these restriction enzymes called? Describe their role in the formation of rDNA.
