Advertisements
Advertisements
प्रश्न
Name the enzymes that are used for the isolation of DNA from bacterial and fungal cells for recombinant DNA technology.
Advertisements
उत्तर
Lysozyme is used for the isolation of DNA from bacterial cells, chitinase is used for the isolation of DNA from fungal cells and cellulase is used for the isolation of DNA from plant cells.
APPEARS IN
संबंधित प्रश्न
Why is the enzyme cellulase needed for isolating genetic material from plant cells and not form the animal cells?
Suggest a technique to a researcher who needs to separate fragments of DNA.
Make a chart (with diagrammatic representation) showing a restriction enzyme, the substrate DNA on which it acts, the site at which it cuts DNA and the product it produces.
Do eukaryotic cells have restriction endonucleases? Justify your answer.
Collect 5 examples of palindromic DNA sequences. Better try to create a palindromic sequence by following base-pair rules.
Answer the following question.
Explain the significance of palindromic nucleotide sequence in the formation of recombinant DNA.
Answer the following question.
Write the use of restriction endonuclease in the formation of recombinant DNA.
There is a restriction endonudease called as EcoRI. What does co part in it stands for?
Which of the given statements is correct in the context of visualizing DNA molecules separated by agarose gel electrophoresis?
'Restriction' in Restriction enzyme refers to ______.
While isolating DNA from bacteria, which of the following enzymes is not required?
The role of DNA ligase in the construction of a recombinant DNA molecule is ______.
Which of the following statements does not hold true for restriction enzyme?
What does H in’ ‘d’ and ‘III’ refer to in the enzyme Hind III?
Given below is the stepwise schematic representation of the process of electrophoresis. Identify the 'alphabets' representing
- Anode end
- smallest/lightest DNA strand in the matrix
- Agarose gel
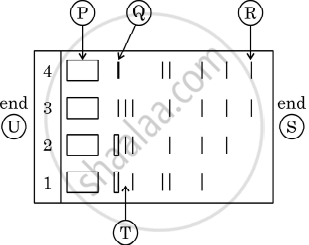
What is elution?
'EcoRI' has played a very significant role in rDNA technology.
- Explain the convention for naming EcoRI.
- Write the recognition site and the cleavage sites of this restriction endonuclease.
State the principle involved in separation of DNA fragments using gel electrophoresis.
How are DNA fragments visualised once they are separated by gel electrophoresis?
