Definition: Histone Octamer
A structural unit composed of eight histone protein molecules around which DNA is wrapped is called histone octamer.
Transfection is the process of inserting a vector into eukaryotic cells.
Definition: Nucleoid
Nucleoid is the region in prokaryotic cells where DNA is organized and associated with proteins, despite the absence of a true nucleus.
Definition: NHC Proteins
Proteins other than histones that are associated with chromatin and help in higher-order DNA packaging and regulation are called non-histone chromosomal (NHC) proteins.
Definition: Nucleosome
The basic repeating unit of chromatin formed by DNA wrapped around a histone octamer is called nucleosome.
Definition: Chromatin
The thread-like complex of DNA and proteins present in the nucleus of eukaryotic cells is called chromatin.
Definition: DNA packaging
The process by which a very long DNA molecule is compactly organised inside the cell nucleus so that it fits within the limited nuclear space and remains functional is called DNA packaging.
Definition: Histones
Positively charged basic proteins rich in lysine and arginine that associate with DNA to help in its packing in eukaryotic cells are called histones.
Definition: Autocatalytic Function
When DNA directs the synthesis of DNA itself, then such function of DNA is called autocatalytic function. Eg. Replication.
Definition: Replication
The process by which DNA duplicates itself is called replication.
Definition: Heterocatalytic Function
When DNA directs the synthesis of chemical molecules other than itself, then such functions of DNA are called heterocatalytic functions. Eg, Synthesis of RNA (transcription), synthesis of protein (Translation), etc.
Definition: Semi-Conservative Replication
Semi-conservative replication is a mode of DNA replication in which each daughter DNA molecule consists of one parental (old) strand and one newly synthesized strand.
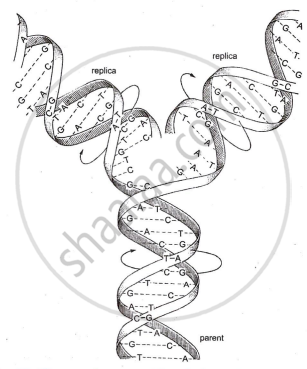
Semi-Conservative Replication
The movement of the ribosome from one end of the mRNA to the other end by the distance of one triplet codon during translation is known as translocation.
A sudden change that occurs in the nucleotide sequence of a gene, causing either a minor or considerable change in the characters of an individual is known as mutation.
Definition: Central Dogma
Central dogma is the principle that genetic information flows in one direction in a cell, from DNA to RNA to protein.
or
Central Dogma is the process by which genetic information flows from DNA to RNA to protein, controlling cellular functions and body structure.
A sequence of three adjacent nucleotides in mRNA that codes for one amino acid is known as a codon.
Translation is the process by which tRNA having anticodon to the codon on the mRNA, supplies amino acids, as per the message on mRNA.
Definition: Transcription
The process of synthesising mRNA from the complementary nucleotide sequence of one strand of DNA, in which uracil replaces thymine, is called transcription.
or
The process of copying genetic information from one strand of the DNA into RNA is termed as transcription.
Definition: Genetic Code
The genetic code is the specific sequence of nitrogenous bases in DNA that determines the order of amino acids in a protein.
Definition: Triplet Codon
A sequence of three nucleotides on mRNA that codes for a specific amino acid is called a triplet codon.
Definition: Translation
The process of protein synthesis in which the message on mRNA is decoded with the help of tRNA to form a specific sequence of amino acids is called translation.
Definition: DNA Fingerprinting
The technique of identifying an individual by analyzing the unique DNA sequence present in each person, similar to fingerprints, is called DNA fingerprinting.
Definition: Operon
A long segment of DNA that contains an operator, promoter and a group of structural genes working together is called an operon.
Definition: Operator
A regulatory DNA sequence that controls transcription by allowing or blocking RNA polymerase binding is called an operator.
Definition: Polycistronic mRNA
A single mRNA molecule that carries information for more than one cistron is called polycistronic mRNA.
Definition: Repressor Protein
A protein that binds to the operator and prevents transcription of structural genes is called a repressor protein.
Definition: Cistron
An individual gene that codes for a single polypeptide chain is called a cistron.
Definition: Regulator Gene
A gene that produces the repressor protein is called a regulator gene.
Key Points: Deoxyribonucleic Acid (DNA)
- DNA was established as the primary genetic material and formally modeled as a double helix by Watson and Crick in 1953.
- The structural blueprint relied heavily on Erwin Chargaff’s chemical base equivalence rules and Rosalind Franklin’s X-ray diffraction data.
- The fundamental building block of DNA is a nucleotide, which comprises a five-carbon deoxyribose sugar, a phosphate group, and a nitrogenous base.
- A nucleotide is distinct from a nucleoside, as a nucleoside contains only the nitrogenous base and pentose sugar without the attaching phosphate group.
- The double helix is formed by two antiparallel polynucleotide chains running in opposite directions (5′→3′ and 3′→5′) coiled in a clockwise, right-handed fashion.
- The physical architecture places the hydrophilic sugar-phosphate backbone on the exterior, while the information-carrying nitrogenous bases stack flat on the interior.
- Structural stability is maintained horizontally by complementary base pairing (A=T via two hydrogen bonds; G=C via three hydrogen bonds) and vertically by strong covalent phosphodiester bonds.
Key Points: Griffith’s Experiment
- Frederick Griffith's 1928 experiments on Streptococcus pneumoniae shifted from developing a pneumonia vaccine to investigating the transmission of bacterial virulence.
- The study compared two distinct bacterial variants: the virulent, encapsulated S (Smooth) strain and the harmless, non-encapsulated R (Rough) strain.
- Baseline experimental controls established that mice survived injections of either the live R strain or the heat-killed S strain independently, but perished when injected with the live S strain.
- The pivotal final experiment revealed that injecting a mixture of live R strain and heat-killed S strain unexpectedly caused fatal pneumonia, resulting in the recovery of live S strain bacteria.
- Griffith concluded that a heritable "Transforming Principle" transferred from the dead S strain and assimilated by the live R strain, converting it into a virulent phenotype.
- While the experiment successfully demonstrated genetic transformation, it was limited by its inability to biochemically identify the transforming substance or confirm it as DNA.
Key Points: Avery, McCarty and MacLeod’s Experiment
- In 1944, Oswald T. Avery, Colin M. MacLeod, and Maclyn McCarty proved that DNA is the genetic material (transforming principle).
- They used cell-free extracts of heat-killed S strain bacteria and mixed them with harmless R strain bacteria.
- Only DNA was able to transform the R strain into the virulent S strain, showing its role in heredity.
- Treatment with proteases and RNases did not stop transformation, proving that protein and RNA are not genetic material.
- Treatment with DNase stopped transformation, confirming that DNA is responsible for it.
- This experiment provided strong evidence that DNA is the hereditary material, though final confirmation came later from the Hershey and Chase experiment.
Key Points: The Hershey-Chase Experiment
- In 1952, Alfred Hershey and Martha Chase proved that DNA is the genetic material using bacteriophages and E. coli bacteria.
- They used radioactive isotopes: ³²P to label DNA and ³⁵S to label proteins.
- Viruses grown in ³²P medium had radioactive DNA, while those grown in ³⁵S medium had radioactive protein.
- These labelled viruses were allowed to infect E. coli, and then blending and centrifugation were done to separate viral coats.
- Only bacteria infected with ³²P-labelled viruses became radioactive, showing that DNA entered the bacterial cells.
- Bacteria infected with ³⁵S-labelled viruses were not radioactive, proving proteins did not enter the cells; hence, DNA is the genetic material.
Key Points: Packaging of DNA Helix
Prokaryote vs Eukaryote Packaging
| Feature |
Prokaryotes |
Eukaryotes |
| Nucleus |
Absent (nucleoid region) |
Present (true nucleus) |
| DNA nature |
Circular, naked (no histones) |
Linear, associated with histones |
| Packaging proteins |
HU proteins, DNA gyrase, Topo I, RNA connectors |
Histones (H1, H2A, H2B, H3, H4) + NHC proteins |
| Packaging mechanism |
Supercoiling + looping |
Nucleosome → Solenoid → Loops → Chromosome |
| Basic repeating unit |
Loop domain |
Nucleosome |
| Levels of compaction |
2 main levels (loops + supercoils) |
5–6 hierarchical levels |
| Charge of packaging proteins |
Positively charged (HU) |
Positively charged (histones) |
Key Points: DNA Replication
- DNA replication is the process by which a DNA molecule makes exact copies of itself, each parental strand acting as a template for a new complementary strand, giving two identical daughter molecules with one old and one new strand each (semi-conservative).
- It occurs during the S-phase (Synthesis phase) of interphase in the cell cycle.
- Proposed by Watson and Crick (1953) and experimentally confirmed as semi-conservative by Meselson and Stahl (1958) in E. coli.
- Of the three proposed models, semi-conservative was proved correct, while conservative and dispersive were disproved.
- Replication is an autocatalytic function (DNA making DNA), unlike heterocatalytic functions, where DNA directs the synthesis of other molecules, like RNA (transcription) or protein (translation).
Key Points: Meselson and Stahl’s Experiment
- The experiment was performed by Meselson and Stahl in 1958 using E. coli, which divides every 20 minutes and is easy to track across generations.
- Bacteria were grown in heavy nitrogen (¹⁵N) medium, then shifted to light nitrogen (¹⁴N) medium, and their DNA was separated by CsCl density gradient centrifugation.
- After the first replication, a single hybrid band appeared, which ruled out the conservative model.
- After the second replication, one hybrid and one light band appeared, which ruled out the dispersive model.
- These results proved that DNA replication is semi-conservative, where each new DNA molecule has one old strand and one new strand.
Key Points: Mechanism of DNA Replication
- Replication starts at the origin (ori), where helicase unwinds the helix into a replication fork, topoisomerase relieves supercoiling, and SSBPs keep the strands apart.
- Primase lays down a short RNA primer, giving DNA polymerase a free 3′-OH end to start adding nucleotides.
- The leading strand is made continuously (one primer); the lagging strand is made discontinuously as Okazaki fragments (many primers).
- DNA Pol I removes the primers and fills the gaps, and DNA ligase seals the nicks into a continuous strand.
- Termination occurs when forks meet at the Ter site (prokaryotes) or when replicons fuse (eukaryotes).
Key Points: Protein Synthesis
- Protein synthesis is the process by which cells produce proteins, which act as structural components, enzymes, and hormones.
- It involves two main steps: transcription (DNA → RNA) and translation (RNA → protein).
- In transcription, genetic information from DNA is copied into mRNA, where uracil (U) replaces thymine (T).
- Central dogma, proposed by Francis Crick (1958), states that information flows from DNA → RNA → protein.
- In retroviruses, reverse transcription occurs (RNA → DNA), explained by Temin and Baltimore (1970) using RNA-dependent DNA polymerase.
Key Points: Transcription
- Transcription is the process in which genetic information from one strand of DNA (template strand) is copied into RNA with the help of RNA polymerase.
- It occurs in the nucleoid in prokaryotes and in the nucleus in eukaryotes; mRNA then moves to the cytoplasm for translation.
- A transcription unit has three parts: promoter (start site), structural gene, and terminator (stop site).
- The process occurs in three stages: initiation (RNA polymerase binds promoter), elongation (RNA chain is formed), and termination (RNA polymerase detaches).
- Only one DNA strand acts as a template (3′→5′), while the other is the coding strand (5′→3′).
- In eukaryotes, primary RNA (hnRNA) is processed by capping, tailing, and splicing to form mature mRNA.
Key Points: Transcription Unit
| Component |
Location |
Function |
| Promoter |
At the 5′ end of the structural gene |
Provides binding site for RNA polymerase and initiates transcription |
| Structural Gene |
Between promoter and terminator |
Contains genetic information to be transcribed |
| Template Strand |
DNA strand with 3′ → 5′ polarity |
Serves as template for RNA synthesis |
| Coding Strand |
DNA strand with 5′ → 3′ polarity |
Does not code directly; used as reference strand |
| Terminator |
At the 3′ end of the coding strand |
Signals the end of transcription |
Key Points: Genetic Code
- The genetic code is the coded information in the base sequence of DNA/mRNA that determines the amino acid sequence in a protein.
- It is a triplet code - three consecutive bases form one codon, proposed by George Gamow (1954).
- There are 64 codons in total: 61 code for amino acids and 3 are stop codons (UAA, UAG, UGA).
- AUG is the start codon and codes for methionine.
- It was deciphered mainly by Nirenberg, Khorana, and Ochoa (poly-U mRNA showed UUU = phenylalanine).
- It is degenerate (one amino acid can have several codons, usually differing in the third base - the wobble effect) and nearly universal.
- A change in the base sequence alters the amino acid sequence, so the code directly controls protein synthesis.
Key Points: Characteristics of the Genetic Code
- Genetic code is a triplet code, that is, three consecutive bases form one codon and specify one amino acid.
- It has distinct polarity and is always read in the 5’ → 3’ direction.
- The genetic code is non-overlapping, so one base is a part of only one codon.
- It is commaless, which means there is no gap or punctuation between successive codons.
- The genetic code is degenerate, so one amino acid may be coded by more than one codon.
- It is universal or nearly universal because the same codon usually specifies the same amino acid in most organisms.
- It is non-ambiguous, so one codon codes for only one specific amino acid.
- AUG is the initiation codon and also codes for methionine.
- UAA, UAG and UGA are stop codons and do not code for any amino acid.
- Codon is written as 5’ AUG 3’, while anticodon is written as 3’ UAC 5’.
Key Points: Mutations and Genetic Code
- A mutation is a sudden change in the DNA sequence that alters the genotype and provides raw material for evolution.
- It may involve deletion (loss) or insertion/duplication (gain) of a DNA segment.
- A point mutation changes a single base pair — e.g., in sickle cell anaemia, glutamate is replaced by valine in the beta-globin gene.
- A frameshift mutation (insertion/deletion of one or two bases) shifts the reading frame, while adding/removing three bases changes whole codons but keeps the frame intact.
- Analogy "RAM HAS RED CAP": one or two extra letters disturb the triplet grouping, but three extra letters keep it unchanged.
Key Points: tRNA – the Adapter Molecule
- tRNA is the adapter molecule (Crick) that picks up amino acids and matches them to the correct mRNA codon.
- Robert Holley proposed the cloverleaf model (2D); in 3D, tRNA looks like an inverted L.
- It has three arms - DHU (binds aminoacyl-tRNA synthetase), anticodon (pairs with codon), TΨC (binds ribosome) - plus a 3′ CCA end for amino acid attachment.
- The amino acid is attached by aminoacyl-tRNA synthetase in a process called aminoacylation (charging), which needs energy.
- Each amino acid has a specific tRNA, with a special initiator tRNA for translation start; no tRNAs exist for stop codons.
Key Points: Translation
- Translation is the process by which the codon sequence on mRNA is decoded with the help of tRNA at the ribosome to form a specific sequence of amino acids in a protein.
- It needs mRNA (template), tRNA (adapter), ribosome (with A, P, and E sites), amino acids, aminoacyl-tRNA synthetase, ATP/GTP for energy, and Mg²⁺ ions.
- Before translation, amino acids are activated and linked to their specific tRNAs (charging) by aminoacyl-tRNA synthetase.
- Initiation: the small ribosomal subunit binds mRNA at the start codon (AUG), the initiator tRNA carrying methionine attaches, and the large subunit joins to form the initiation complex.
- Elongation: amino acids are added one by one through codon–anticodon pairing, peptide bonds form between them, and the ribosome moves forward by one codon at a time (translocation).
- Termination occurs at a stop codon (UAA, UAG, UGA), where release factors free the polypeptide and the ribosomal subunits separate.
Key Points: Mechanism of Translation
| Stage |
Key Events |
Enzymes / Factors Involved |
| Activation of Amino Acids |
Amino acids are activated by ATP and attached to specific tRNA molecules |
Aminoacyl-tRNA synthetase, ATP, Mg²⁺ |
| Role of Ribosome |
Ribosome provides sites for mRNA binding and peptide synthesis; has A site and P site |
rRNA, ribosomal proteins |
| Initiation |
mRNA binds to small ribosomal subunit; initiator tRNA binds to start codon (AUG) at P site; large subunit joins |
Initiation factors, Mg²⁺, GTP |
| Elongation |
Aminoacyl-tRNA binds to A site; peptide bond forms; ribosome translocates along mRNA |
Peptidyl transferase, elongation factors, GTP |
| Termination |
Stop codon (UAA, UAG, UGA) is reached; polypeptide released; ribosome dissociates |
Release factors, GTP |
| Post-translational Modification |
Polypeptide undergoes folding and chemical modifications |
Deformylase, peptidases |
| Protein Translocation |
Proteins synthesized on free or bound ribosomes are transported to correct cellular locations |
ER membrane, Golgi apparatus |
Key Points: Regulation of Gene Expression
- Gene regulation = switching genes ON or OFF based on the cell's requirement and stage of development.
- In eukaryotes, regulation occurs at 4 levels: Transcriptional (primary transcript), Processing (splicing), Transport (mRNA from nucleus to cytoplasm), and Translational.
- In prokaryotes, control of transcriptional initiation rate is the main site for gene expression control.
- E. coli produces β-galactosidase to break lactose → galactose + glucose. If lactose is absent, the enzyme is not produced, proving the environment regulates gene expression.
- Enzymes synthesized based on substrate availability are called inducible enzymes. The process = induction; the triggering molecule = inducer. This is a positive control.
- Feedback repression = when the end product (e.g., amino acid) is already available, genes for its production are switched OFF. This is a negative control.
Key Points: Operon Concept
- The operon concept (Jacob & Monod, 1961) explains how related genes are regulated together as one unit, based on the lac operon in E. coli.
- An operon has four parts: regulator (i), promoter (p), operator (o), and structural genes (z, y, a) - coding for β-galactosidase, permease, and transacetylase.
- The operator acts as an ON/OFF switch; when active, all three genes are transcribed into a single polycistronic mRNA (each gene = one cistron).
- The repressor (from the regulator gene) controls the operator - bound = OFF, unbound = ON.
- Inducible system (lac operon): the inducer (lactose) inactivates the repressor → transcription ON.
- Repressible system: a co-repressor activates the repressor → transcription OFF.
Key Points: The Lac Operon
- The lac operon is an inducible operon in E. coli, proposed by Jacob and Monod (1961), that controls the metabolism of lactose.
- It consists of a regulator gene (i), promoter (P), operator (O), and three structural genes - lac z, lac y, lac a - coding for β-galactosidase, permease, and transacetylase respectively.
- When lactose is absent: the regulator gene produces an active repressor that binds the operator and blocks RNA polymerase, so the operon remains switched OFF.
- When lactose is present: lactose is converted into allolactose (the inducer), which binds the repressor and inactivates it, leaving the operator free.
- RNA polymerase then transcribes the structural genes into a single polycistronic mRNA, producing the enzymes that break down lactose - the operon is now switched ON.
Key Points: Genomics
- The genome is the complete genetic material of an organism, while genomics is the study of the entire genome, including sequencing, mapping, and gene functions.
- The term genome was introduced by H. Winkler (1920) and genomics by T.H. Roderick (1986).
- Genomics uses DNA sequencing, recombinant DNA technology, and bioinformatics tools to study the structure and function of genes.
- Genomics is divided into structural genomics (mapping and sequencing of genomes) and functional genomics (study of gene expression and function).
- Applications of genomics include the development of transgenic crops, gene therapy, forensic analysis using genetic markers, and the production of useful proteins and biofuels in microbes.
Key Points: Human Genome Project
- The Human Genome Project (HGP) was an international mega-project launched in 1990 and completed in 2003, coordinated mainly by the U.S. DOE and NIH.
- Its aim was to identify all human genes and sequence the entire human genome of about 3 billion base pairs.
- The main goals were to identify genes, sequence the genome, store the data, develop analysis tools, transfer technologies, and address ethical issues.
- Methodology: DNA was isolated, fragmented, cloned into vectors like BACs and YACs, sequenced by automated methods, and assembled using computers.
- Salient features: the genome has ~3 billion base pairs and 20,000–25,000 genes; less than 2% codes for proteins, and humans are 99.9% identical.
- Most genetic variation between individuals is due to SNPs (single-nucleotide polymorphisms).
- Applications: disease gene mapping, early diagnosis, personalised medicine, evolutionary studies, and advances in biotechnology.
- ELSI (Ethical, Legal, Social Issues): genome data must be kept confidential to prevent misuse and discrimination.
Key Points: DNA Fingerprinting
- DNA fingerprinting is a technique used to identify an individual by analysing the unique DNA pattern present in every person (except identical twins).
- It is based on satellite DNA, especially VNTRs (Variable Number Tandem Repeats) - short sequences repeated in tandem, whose number varies among individuals and creates DNA polymorphism.
- Principle: the differences in VNTR repeat number produce DNA fragments of different lengths, which appear as a unique banding pattern.
- Steps: DNA isolation → PCR amplification → restriction digestion → gel electrophoresis → Southern blotting → probe hybridisation → autoradiography → comparison of band patterns.
- Applications: forensic identification, paternity/maternity testing, pedigree studies, medical research, conservation biology, and evolutionary/anthropological studies.
Key Points: Packaging in Prokaryotes
- Prokaryotes like E. coli do not have a true nucleus. Their DNA is present in a region called nucleoid.
- The DNA is very long but the cell is very small, so the DNA must be tightly packed inside the cell.
- The DNA becomes circular and folded into loops. About 40–50 loops are formed to reduce its size.
- The DNA is further coiled and supercoiled with the help of HU proteins, DNA gyrase and topoisomerase, so it fits inside the cell.
Key Points: Regulation of gene expression
- Gene expression is a multistep process by which a gene is regulated and its product (polypeptide/protein) is formed.
- In eukaryotes, gene expression is regulated at different levels such as transcription, RNA processing, mRNA transport, and translation.
- Genes are expressed only when needed to perform specific functions, for example β-galactosidase enzyme in E. coli helps in breaking lactose into glucose and galactose.
- If lactose is absent, E. coli does not produce β-galactosidase, showing that gene expression depends on environmental and metabolic conditions.
- Some bacterial genes are inducible, meaning they are switched on only in the presence of a substrate; this process is called induction and works by positive control.
Key Points: Genomics
- The complete genetic constitution or one complete set of chromosomes of an organism is called the genome, and its study is known as genomics.
- Genomics involves sequencing, mapping, analysis of genes, and understanding their functions.
- Genomics is of two types: structural genomics (mapping and sequencing of genomes) and functional genomics (study of gene functions and expression).
- Comparative genomics is done by sequencing organisms like yeast, fruit fly and mouse to understand human genes.
- Genomics has applications in medicine, agriculture, biotechnology and forensics, including gene therapy, transgenic crops and production of useful proteins.
