Advertisements
Advertisements
Question
Give the steps in PCR or polymerase chain reaction with suitable diagrams.
Advertisements
Solution
Polymerase Chain Reaction (PCR) is the process of in vitro amplification of the gene of interest using a PCR machine.
i. PCR can generate a billion copies of the desired segment of DNA or RNA, with high accuracy and specificity, in a few hours.
ii. The process of PCR is completely automated and involves automatic thermal cycles for denaturation and renaturation of double-stranded DNA.
iii. The device required for PCR is called a thermal cycler.
iv. Requirements for polymerase chain reaction:
- DNA containing the desired segment to be amplified
- several molecules of four deoxyribonuclueoside triphosphates (dNTPs)
- excess of two primer molecules
- heat-stable DNA polymerase and
- appropriate quantities of Mg++ ions.
Mechanism of PCR:
At the start of PCR, all the requirements are mixed together in ‘Eppendorf tube’ and the following operations are performed sequentially:
Step i: Denaturation
The reaction mixture is heated to a temperature (90–98oC) to separate two strands of desired DNA. This is called denaturation.
Step ii: Annealing
The mixture is allowed to cool (40–60oC) that permits the pairing of the primer to the complementary sequences in DNA. This step is called annealing.
Step iii: Primer extension / Polymerization
The temperature (70–75°C) allows thermostable Taq DNA polymerase to use single-stranded DNA as a template and adds nucleotides. This is called primer extension. It takes around two minutes duration.
v. One cycle takes around 3 to 4 minutes.
vi. To begin the second cycle, DNA is again heated to convert double-stranded DNA into single strands.
vii. In an automatic thermal cycler, the above three steps are automatically repeated 20-30 times. Thus, at the end of ‘n’ cycles, 2n copies of DNA segments are produced.
viii. The machine performs the entire operations automatically and precisely.
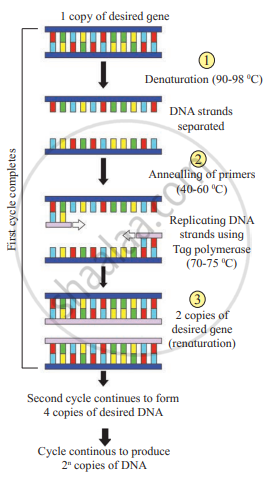
DNA replication through a polymerase chain reaction.
APPEARS IN
RELATED QUESTIONS
Give an account of the Blue-White Method of selection of recombinants.
Why are bio-fertilizers preferred over chemical fertilizers?
Enlist and write in brief about the different biological tools required in rDNA technology.
The technique which involves addition or deletion of genes is ______.
What is gene cloning? Explain different tools used for it.
The time taken to complete one cycle of PCR is around ______.
Identify the process by which bacteria protect their own DNA from the attack of restriction enzymes.
______ is the soil bacterium that causes crown gall characterized by tumors in plants.
The transfer of genetic material from one bacterium to another through the mediation of a viral vector is termed as ______.
The restriction enzymes used as molecular scissors are type of ______.
